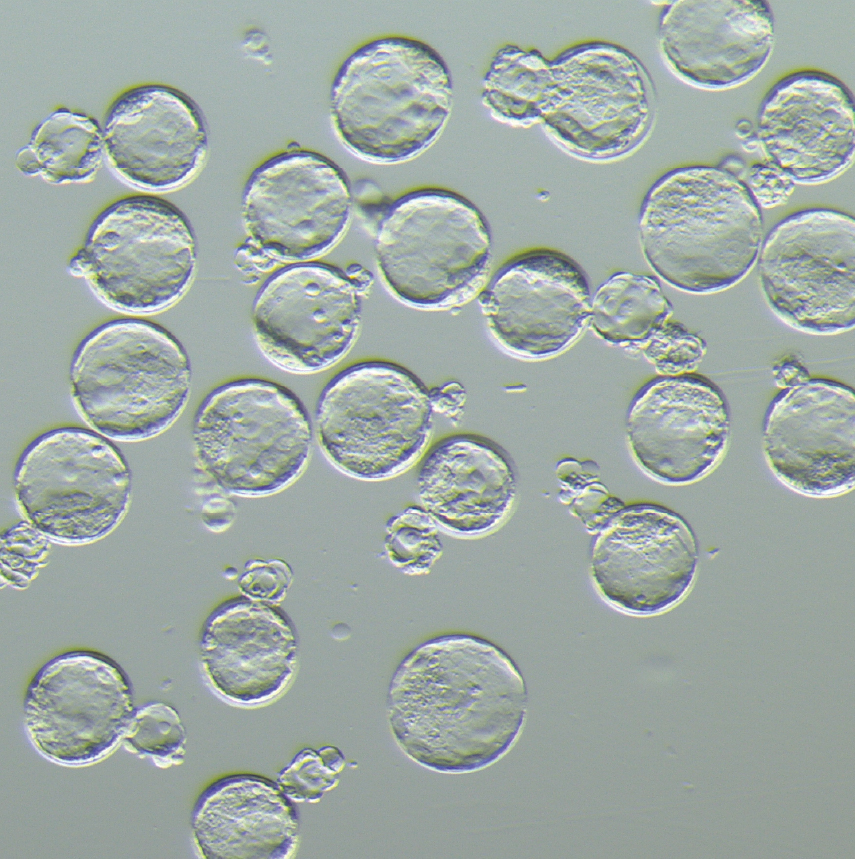
The life of mammals starts with fertilization of an egg, which subsequently undergoes multiple rounds of cell division and develops into a complete organism. During mammalian pre-implantation, transcription factors (TFs) play a crucial role in activating the zygotic genome and determining lineage segregation into the inner cell mass and trophectoderm, which give rise to the embryo proper and extraembryonic tissue (e.g. placenta). Notably, specialized TFs, called pioneer factors, have unique abilities that bind nucleosomal DNA and modulate epigenetic and chromatin states by facilitating the recruitment of chromatin remodelers and histone modifiers. We study how pioneer factors act as top regulators of each cell-stage regulatory network and define cell potential during developmental transitions.
Mechanisms of epigenome regulation by pioneer factors in the early embryo
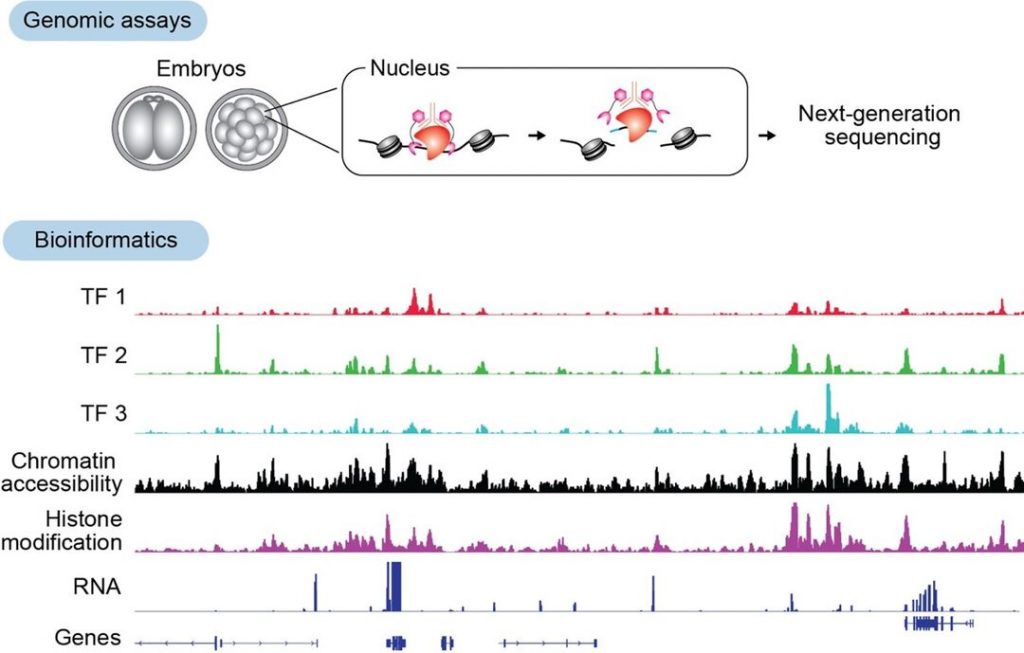
Recent advances in low-input genomics technologies have enabled the identification of genome-wide protein-DNA interaction regions in mammalian embryos. We have adapted and applied the CUT&Run, CUT&Tag, and TIP-seq methods for mouse embryos with low-input numbers (Kobayashi et al., Development 2026; Kobayashi and Tachibana, Methods Mol Biol. 2025; Gassler and Kobayashi et al., Science 2022). By combining these approaches with knockdown and knockout strategies, we aim to establish how TFs regulate epigenetic states and transcription to determine cell fate.
In parallel, we employ a proteomics approach using state-of-the-art Orbitrap Astral mass spectrometry at University of Dundee to dissect how transcription factors switch between activating and repressive functions. Through integrating genomics, proteomics, and functional perturbation, we aim to uncover the molecular mechanisms by which pioneer factors orchestrate epigenetic reprogramming in the early embryo.
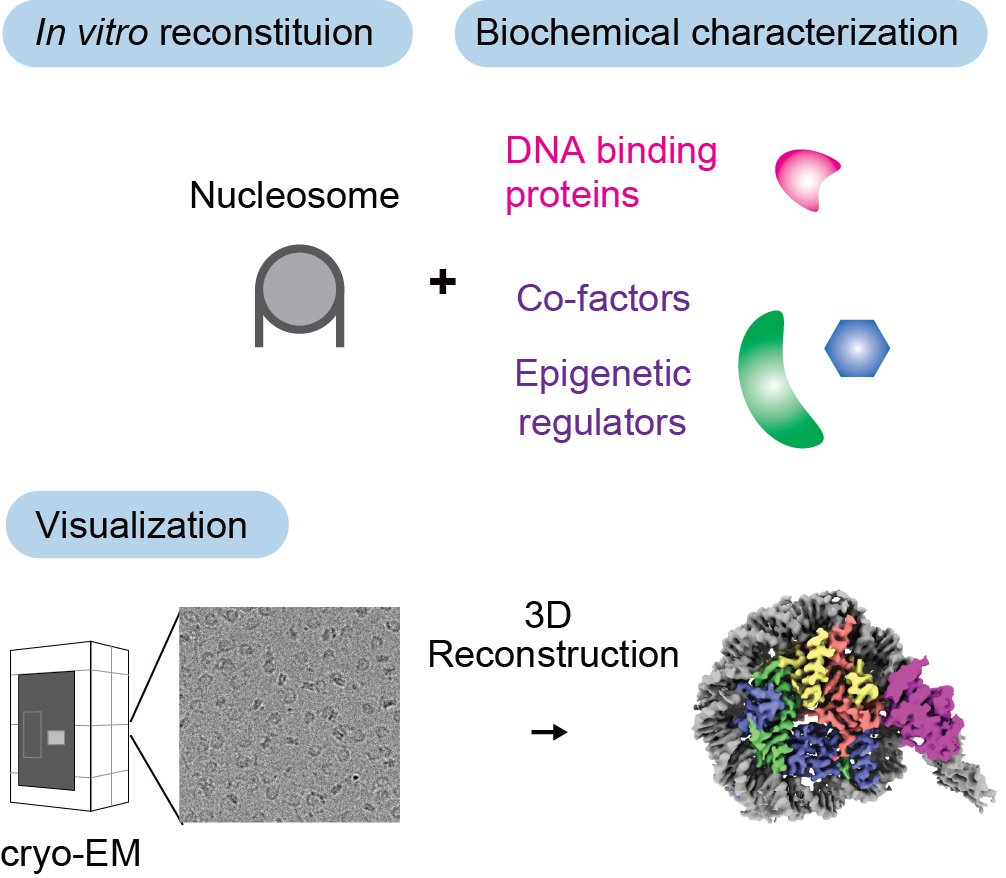
In vitro reconstitution and visualization of chromatin
To uncover the molecular basis by which TFs control chromatin and epigenome, we take an in vitro reconstitution approach. We purify recombinant proteins and assemble defined complexes with nucleosomes in a test tube, allowing us to characterize their properties by biochemical assays. Using cryo-Electron Microscopy (cryo-EM), we further visualize these complexes and determine their structural basis at the molecular and atomic level (Kobayashi et al., Nat Struct Mol Biol 2024; Kobayashi et al., Commun Biol 2019). We have access to a state-of-the-art Glacios, 200kV cryogenic transmission electron microscope for screening and dataset collection.